cdkbook
Molecular Descriptors
The previous chapter discussed prediction of molecular properties. Understanding them is a core aspect of chemistry. Computational tools help here. Quantum Chemistry promises to calculate many molecular properties accurately from mere knowledge of atom locations and their electronic properties, but suffers from a badly scaling algorithm. Instead, approaches using the chemical graph as a core representation are less accurate, but much more faster.
Quantitative Structure-Activity Relationship (QSAR) modeling uses this latter approach to predict molecular properties. QSAR approaches commonly use molecular descriptors which numerically describe molecular allowing statistical methods to correlate these descriptors to end points, such as the logP property. A far more detailed explanation can be found in [1].
This chapter will discuss the descriptor API present in the CDK to calculate molecular descriptors. It should be noted that the CDK also contains atom, atom pair, bond, and protein descriptors, using a similar API.
Descriptors and Specifications
The CDK makes a distinction between descriptor specifications and their implementations. The reason is simple: the CDK has implementations of descriptor specifications. Such an implementation may depend on data available in the CDK, such as isotopic masses which get more accurate over time. Consequently, the calculated properties will deviate when this data gets updated: the descriptor implementation changes even when the algorithm stays the same.
Practically, things are even a bit more complicated, as the CDK is different in another way. The CDK project decided it was better to have parameterizable implementations: one descriptor instance that can calculate the number of carbons, but also the number of nitrogens. After all, we do not want too much code replication.
Therefore, the CDK has two core interfaces: IDescriptor and
IImplementationSpecification [2]. The first is an actual
descriptor instance,
parameterized to calculate particular output. The second links this
descriptor to a formal specification, outlined in the Blue Obelisk
Descriptor Ontology [3,4].
IImplementationSpecification
Let’s first look at the IImplementationSpecification interface.
This specification interface provides provenance information that allows you to keep
track how your descriptor values were calculated. Of course, you may
simply ignore that whenever you do not need that information.
This interfaces defines four fields:
- specification reference
- implementation title
- implementation identifier
- implementation vendor
This information links the CDK descriptor implementation to a formally defined descriptor algorithm, pointed to by the specification reference. The other three fields provide information on the CDK implementation, allowing one to mix descriptor calculation by various tools and to keep track of what value came from what vendor.
For example, we can inspect these field values for the TPSA descriptor (see Section 18.3):
Script 17.1 code/TPSASpecs.groovy
descriptor = new TPSADescriptor()
specs = descriptor.specification
println "Title: " + specs.implementationTitle
println "Reference: " + specs.specificationReference
println "Vendor: " + specs.implementationVendor
println "Identifier: " + specs.implementationIdentifier
This code provides us with these details:
Title: org.openscience.cdk.qsar.descriptors.molecular.TPSADescriptor
Reference: http://www.blueobelisk.org/ontologies/chemoinformatics-algorithms/#...
tpsa
Vendor: The Chemistry Development Kit
Identifier: 2.11
The identifier values originally referred to a String which held the source code repository’s commit number. But ever since we moved from a Subversion source code repository to Git, we no longer had commit numbers, but commit hashes again. But both identifiers have the function that they identified a specific revision of the implementation allowing people to accurately recalculate that descriptor value, given the same input. Well, no one seems to worry about that much, but it is with this framework possible anyway.
IDescriptor
As the previous code example (Script ??) shows, the
specification is for an descriptor implementation and we used
the getSpecification() (.specifiction in Groovy)
to get the specification information. This method is defined by
the IDescriptor interface.
The interface introduces the following additional methods:
getDescriptorNames()getParameterNames()getParameterType(String name)setParameters(Object[] params)getParameters()
The CDK descriptor API differs from other tools in that it structures descriptors around the algorithms that calculate them. Therefore, each descriptor returns one or more descriptor values. The first methods lists the descriptor value names, and it should be noted that those are specifically for the used parameters, as we will see later. The default descriptor value names are listed in Appendix C.5.
The next two methods, getParameterNames() and getParameterType(String name),
allow us to discover which parameters a particular descriptor has, if any. For example,
we can check those of the AtomCountDescriptor:
Script 17.2 code/AtomCountDescriptorParams.groovy
descriptor = new AtomCountDescriptor()
descriptor.parameterNames.each { name ->
type = descriptor.getParameterType(name).class.name
println "$name -> $type"
}
which tells us how we can tune the descriptor calculation (see also Section 19.5):
elementName -> java.lang.String
The last two methods allow is to change the current parameter values
(setParameters(Object[] params)), and what those active values are
(getParameters()). These values are in the same order as the parameter
names. For example, we can see if the AromaticAtomsCountDescriptor calculates
aromaticity itself by default:
Script 17.3 code/AromaticAtomCountDescriptorParams.groovy
descriptor = new AromaticAtomsCountDescriptor()
println "Values:"
descriptor.parameters.each { param ->
type = param.class.name
println "$type -> $param"
}
println "Updating the value..."
Object[] newValues = [ Boolean.TRUE ]
descriptor.setParameters(newValues)
println "New values:"
descriptor.parameters.each { param ->
type = param.class.name
println "$type -> $param"
}
Note that I used the Groovy syntax to create an array here, because the Java syntax overlaps with the Groovy syntax for specifying closures [5]. This code shows us:
Descriptor names:
checkAromaticity -> java.lang.Boolean
Values:
java.lang.Boolean -> false
Updating the value...
New values:
java.lang.Boolean -> true
IMolecularDescriptor
With this information about the descriptor now clear, it is time to look at a descriptor
value calculation. A molecular descriptor in the CDK is symbolized by the
IMolecularDescriptor interface, which extends the IDescriptor interface,
as shown in Figure 19.1.
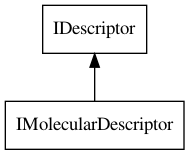
Figure 19.1: The IDescriptor interface has a few derived interfaces, but only IMolecularDescriptor is shown here.
The relevant method now is the calculate(IAtomContainer) method, which returns
a DescriptorValue. This class is returned rather than a double, because descriptors
can calculate multiple types. The most common two types are a single Double and
a Double[], followed by the integer variants. However, a descriptor could also
return a cardinal value, and a boolean. But before we go into the details of the
actually calculated descriptor values, let’s look at TPSA calculation in a bit more detail
then we did in Script ??:
Script 17.4 code/TPSACalc.groovy
descriptor = new TPSADescriptor()
result = descriptor.calculate(oxazone)
println "Specification: " + result.specification
println "Parameters names: " + result.parameterNames
println "Parameters values: " + result.parameters
println "Exception: " + result.exception
println "Names: " + result.names
value = result.getValue()
The output shows us that quite some metadata is preserved:
Specification: org.openscience.cdk.qsar.DescriptorSpecification@5fe5c68b
Parameters names: [checkAromaticity]
Parameters values: [false]
Exception: null
Names: [TopoPSA]
The descriptor specification is kept for reference, as well as the parameters, and the descriptor value names. This is all the information you need to report in a publication so that other people can recalculate the exact values you calculated.
Additionally, if the calculation failed (because the input structure had error, had missing information, like no 3D coordinates), then an exception is stored and returned too:
Script 17.5 code/DescriptorCalcException.groovy
descriptor = new MomentOfInertiaDescriptor()
result = descriptor.calculate(oxazone)
exception = result.exception
println "Exception:\n" + exception
The moment of inertia descriptor requires 3D coordinates, which are not provided in the above script. Therefore, we get an exception:
Exception:
org.openscience.cdk.exception.CDKException: Molecule must have 3D coordinates
IDescriptorResult
With all this context described, it is time to look at the actual calculated values. It was already noted that a molecular descriptor implementation in the CDK returns one or more values, each of which can be of a varying type. Indeed, if we look at Appendix C.5 we see that some descriptors return one numerical value, while others return many values. It should also be noted, that depending on parameter values set, the actual number of calculated numbers can vary!
Script 17.6 code/DescriptorResultLength.groovy
descriptor = new MomentOfInertiaDescriptor()
result = descriptor.calculate(methane)
value = result.value
println "Calculated values: " + value.length()
Which shows us that the molecular inertia descriptor calculates more than one value:
Calculated values: 7
The IDescriptorResult interface is currently implemented by various classes,
outlined in Figure 19.2. Each of the classes has a slightly different
API to get the actual values. The IntegerResult, DoubleResult,
and BooleanResult classes have the methods intValue(),
doubleValue(), and booleanValue() respectively.
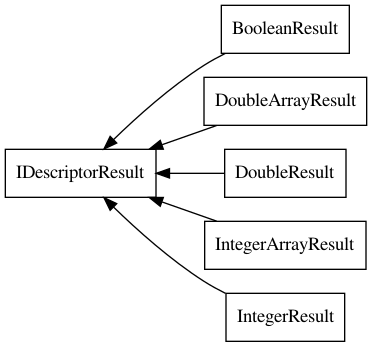
Figure 19.2: The IDescriptorResults interface has several implementation, each wrapping calculated descriptor values.
The two array variants, IntegerArrayResult and DoubleArrayResult,
work slightly different, and both provide a get(int) method to iterate over all
values. For example, for the molecular inertia descriptor values for methane it would
look like:
Script 17.7 code/DoubleArrayGetValue.groovy
value = descriptor.calculate(methane).value
for (i in 0..(value.length()-1)) {
println i + " " + value.get(i)
}
This returns us all inertia values, and note that the index starts at zero:
0 3.1955750763324886
1 3.1945914391012566
2 3.1941764686200322
3 1.0003079070516474
4 1.0004378617544136
5 1.0001299147011136
6 1.1190669469245764
Counting Nitrogens and Oxygens
Now that we know how the general API works, we can calculate custom descriptors, say the
nitrogenCount and oxygenCount. We reuse the AtomCountDescriptor for that, and
set the parameter:
Script 17.8 code/SpecificAtomCountDescriptor.groovy
descriptor = new AtomCountDescriptor()
Object[] params = [ "N" ]
descriptor.setParameters(params)
calculated = descriptor.calculate(ethanol)
nitrogenCount = calculated.value
label = calculated.names[0]
println "Number of nitrogens ($label): $nitrogenCount"
params = [ "O" ]
descriptor.setParameters(params)
calculated = descriptor.calculate(ethanol)
oxygenCount = calculated.value
label = calculated.names[0]
println "Number of oxygens ($label): $oxygenCount"
And this returns us the nitrogen and oxygen counts:
Number of nitrogens (nN): 0
Number of oxygens (nO): 1
It should be noted too, that the descriptor labels, given in brackets in the output, have been updated according to the change in descriptor parameters. This is not necessarily the case for all descriptors, but many take this approach.
References
- Wikberg J, Eklund M, Willighagen E, Spjuth O, Lapins M, Engkvist O, et al. Introduction to Pharmaceutical Bioinformatics. 2018.
- Steinbeck C, Hoppe C, Kuhn S, Floris M, Guha R, Willighagen E. Recent Developments of the Chemistry Development Kit (CDK) - An Open-Source Java Library for Chemo- and Bioinformatics. Curr Pharm Des [Internet]. 2006 Jun 1;12(17):2111–20. Available from: https://cdk.github.io/cdk-paper-2/ doi:10.2174/138161206777585274 (Scholia)
- Guha R, Howard MT, Hutchison GR, Murray-Rust P, Rzepa HS, Steinbeck C, et al. The Blue Obelisk-interoperability in chemical informatics. JCIM. 2006 Feb 22;46(3):991–8. doi:10.1021/CI050400B (Scholia)
- Spjuth O, Willighagen E, Guha R, Eklund M, Wikberg J. Towards interoperable and reproducible QSAR analyses: Exchange of datasets. J Cheminform. 2010;2(1):5. doi:10.1186/1758-2946-2-5 (Scholia)
- https://stackoverflow.com/questions/9363550/how-to-prevent-getting-a-groovy-boolean-in-an-object-array